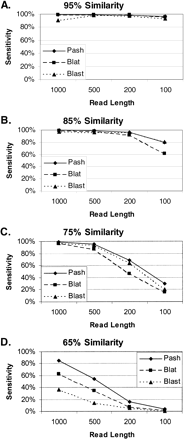
Simulated anchoring. Randomly selected sequence fragments of varying lengths (1000, 500, 200, or 100 bp) from the human genome were randomly mutated to be 95% (A), 85% (B), 75% (C), or 65% (D) identical to their original sequence. The fraction of fragments that were correctly mapped to their original location by Pash, BLAT, and BLAST indicate the sensitivity of these methods at varying levels of sequence identity.











