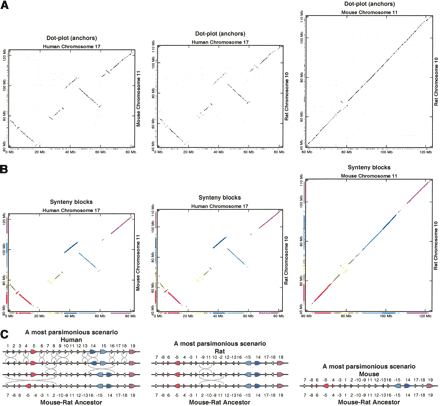
Region on human chromosome 17, mouse chromosome 11, rat chromosome 10. (A) Local two-way similarities produced by PatternHunter (darker ones are longer; some short mouse–rat ones were removed for legibility). Synteny blocks appear as ±45° diagonals. Repeats tend to appear as discrete grids with irregular spacing. (B) After removing repeated regions and combining close alignments, GRIMM-Synteny computes 19 large-scale synteny blocks (at least 300 kb). The same synteny blocks are shown in each pair of species, using consistent colors. (C) The arrangement (order and orientation) of the 19 synteny blocks in this region is shown for each genome. MGR determined that there is a unique median ancestor associated with the most parsimonious evolutionary scenarios of the three genomes in this region; note that it coincides with mouse. The arrangement of blocks in this region implies there were at least 12 inversions between human and the median, and at least two inversions between rat and the median. The chronological order of the inversions cannot be inferred from this method. Note also that the minimum number of inversions required to convert the human block order in this region into the rat order is 14, so the solution shown is optimal, and all optimal solutions have equal block arrangements on median and mouse.











