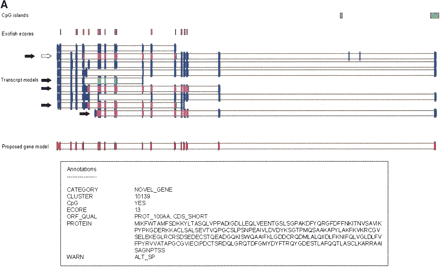
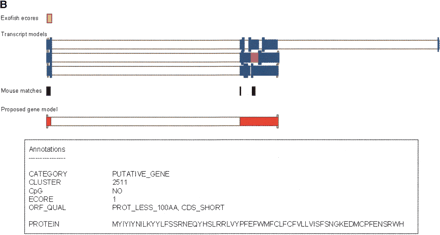
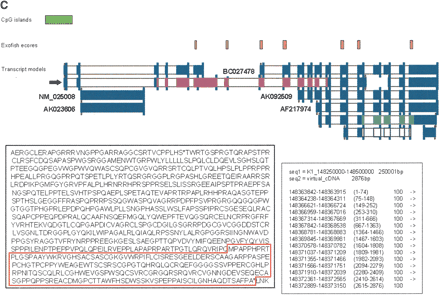
(A) Example of a new gene supported by the CNSLT cDNA resource. Human curation of the identified gene models was performed by using a graphical interface. The transcript models of cDNA clones are represented by blue bars, whereas the model proposed for the gene is represented at the bottom of the figure by red bars (for a detailed explanation, see Results). An empty arrow indicates the CNSLT cDNA clone used for the construction of the proposed gene model. Filled arrows indicate the cDNA clones assembled on the genome. CpG islands are represented by green boxes; human-Tetraodon ecores, by orange boxes. Coding regions with an in-frame stop codon upstream of an initiator ATG are represented by magenta bars. When such stop codons could not be identified, the coding regions are represented by pale turquoise bars. Annotations found for this proposed gene model are listed in the boxed part of the figure. PROT_100AA indicates CDS of at least 300 bp; CDS_SHORT, coding region spanning <50% of the model sequence; and ALT_SP, alternative splicing. (B) Example of a putative gene. The transcript models of cDNA clones are represented by blue bars, whereas the model proposed for the gene is represented at the bottom of the figure by red bars (for a detailed explanation, see Results). A human–Tetraodon ecore, which supports the first exon of the proposed gene model, is represented by an orange box. A coding region of 64 amino acids with an in-frame stop codon upstream of an initiator ATG is represented by magenta bars. Matches to the mouse genome are indicated by black bars. Annotations found for this proposed gene model are listed in the boxed part of the figure. PROT_LESS_100AA indicates CDS <300 bp. (C) Example of an extension of an already annotated gene, using the CNSLT cDNA resource. The transcript models of cDNA clones or RefSeq and GenBank transcripts are represented by filled blue bars. CpG islands are represented by green boxes, and human–Tetraodon ecores by yellow boxes. The filled arrow points to the CNSLT cDNA clone extending the annotated gene. (Inset, right) The exons predicted by the alignment of the virtual cDNA sequence and the human genome assembly, using the sim4 algorithm (for detailed explanation, see text). Color code for coding regions: stop-ATG-stop, magenta bars; ATG-stop, pale turquoise bars; and stop-ATG, white bars. (Inset, left) The extension of the CDS from BC027478 (red box) using the CNSLT cDNA resource.











