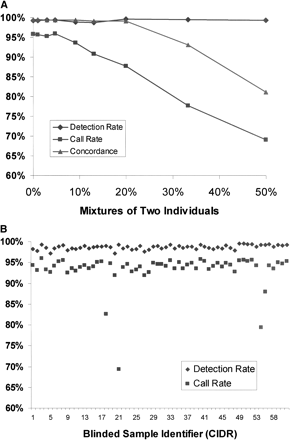
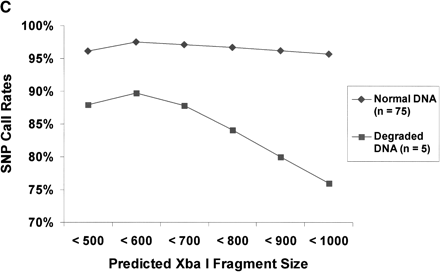
(A) Detection of sample contamination. Sample DNAs from two individuals were progressively mixed together and genotyped as mixtures. The detection rate is a measure of the number of SNPs passing the signal vs. noise discrimination filter. The call rate is the percentage of SNPs that were assigned a genotype instead of a no call. The concordance rate is based on comparison of genotype calls with SBE reference genotypes for one of the two individuals. At 50% mixture, the concordance rate was nearly identical compared to SBE reference genotypes from either individual (data not shown). (B) Blinded samples from the CIDR. Detection and call rates from 61 blinded samples from the CIDR. Blinded samples containing mixtures of two individuals were identified as having high detection rates, but low call rates. Other blinded samples included eight samples that were previously problematic when genotyped at STR loci. The samples also included 18 members of a family affected with chronic mucocutaneous candidiasis and thyroid disease, as well as CEPH pedigree samples. (C) Degraded sample DNA and SNP call rates. Call rates were calculated for each SNP across multiple experiments. There were five experiments in the case of degraded sample DNAs, and for comparison, 75 experiments in the case of normal DNAs. SNPs were binned into six groups on the basis of the predicted lengths of the restriction fragments containing the SNPs. Each bin represented restriction fragment lengths in increments of 100 bp, and SNP call rates were averaged within each of the bins.











