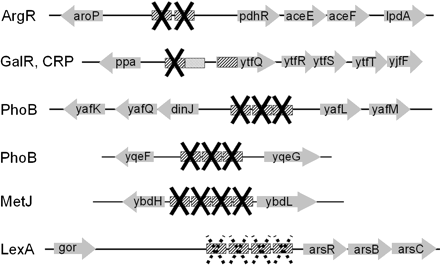
Summary of binding site knockouts. (Solid X) Predicted binding sites that were knocked out. (Dashed X) Knockouts of four predicted LexA-binding sites in the gorR–arsR IGR that were not successful, possibly because of lethality. Distances between genes are approximately to scale. In the uppermost construct, two predicted ArgR-binding sites (cross-hatched boxes) were knocked out in the aroP–pdhR IGR. In the construct shown below that one, only one of the two predicted GalR-binding sites (cross-hatched boxes) in the ppa–ytfQ IGR was knocked out; the predicted CRP site (stippled boxes) was not knocked out. The other constructs created successfully were knockout of three predicted PhoB sites (cross-hatched boxes) in the dinJ–yafL IGR; knockout of three predicted PhoB sites (cross-hatched boxes) in the yqeF–yqeG IGR; knockout of four predicted MetJ sites (cross-hatched boxes) in the ybdH–ybdL IGR. The predicted ArgR-binding sites are 241 bp upstream of aroP and 260 bp upstream of pdhR; the predicted PhoB sites are 103 bp upstream of dinJ and 73 bpupstream of yafL; the predicted GalR and CRP sites are 164 bp upstream of ppa and 129 bpupstream of ytfQ; the predicted MetJ sites are 18 bpupstream of ybdH and 58 bpupstream of ybdL; the predicted PhoB sites are 37 bpupstream of yqeF and 181 bpupstream of yqeG; the predicted LexA sites are 328 bp downstream of gor and 556 bpupstream of arsR (by “upstream,” here we mean the distance between the predicted binding sites and the start codon of the gene).











