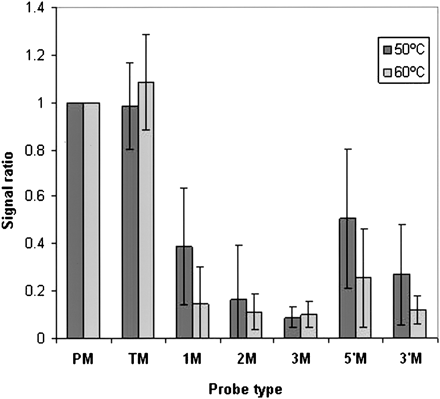
Effect of mismatches within MIRs on microarray signal intensity. Signals from hybridization of 60-mer probes with HeLa lcRNA at either 50 or 60°C were compared. Probes containing MIR at the 5′ end followed by nonhuman genome sequences were examined. Each probe containing a wild-type MIR (PM) was compared with a set of probes with the following mismatches: (TM) A probe with a wild-type MIR and six mismatches in the adjacent nonhuman genome sequence region; (1M) a probe with one mismatch at position 10 of the MIR sequence; (2M) a probe with two mismatches at positions 8 and 17 of the MIR; (3M) a probe with three mismatches at positions 6, 12, and 18 of the MIR. (5′M) a probe with a block of four mismatches at the 5′ end of the MIR; (3′M) a probe with a block of six mismatches at the 3′ end of the MIR. Data are presented as mean signal ratio ± SEM for 42 sets of probes.











