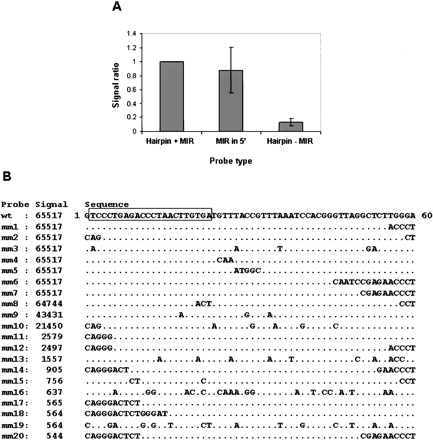
MIRs are responsible for signals from probes of precursor sequences. Signals from hybridization of 60-mer probes with HeLa lcRNA are presented. (A) Three probes were considered for each MIR as follows: (1) precursor sequence containing the full MIR (hairpin + MIR); (2) MIR in the 5′ end of the probe, followed by nonhuman genome sequences (MIR in 5′); (3) Precursor sequence containing no more than 16 nucleotides of the MIR sequence (hairpin–MIR). Only probe sets in which the microarray signal of the hairpin + MIR probe was >2000 were considered. For each probe set, the signal intensities for the three types of probes were calculated as a ratio of the signal intensity of the hairpin + MIR probe. The MIR in 5′ and hairpin–MIR bars show the average computed for 23 sets of probes. (B) A detailed example of a mismatch analysis of pre-MIR-125b. The pre-MIR-125b probe and its respective mismatch (mm) probes from MirChip1 are presented. Wild-type (wt) probe contains MIR-125b at position 2 to 23 (boxed). Probes with mismatches are listed below the wild-type probe with the location of the substituted nucleotides specified. The signal column presents the microarray signal intensity observed for each probe. Similar results were obtained for let-7a and miR-30c.











