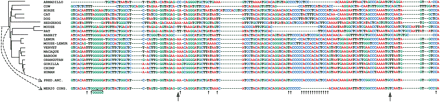
Example of reconstruction of an ancestral Boreoeutherian sequence based on actual orthologous sequences derived from a MER20 retrotransposon. Arrows indicate positions where the reconstructed ancestor differs from the MER20 consensus. Longer arrows indicate the positions where the knowledge of the MER20 consensus sequence would have changed the ancestral base prediction. The position of the human sequence displayed is chr7:115,739,755-115,739,899 (NCBI build 34). The alignment of the flanking nonrepetitive DNA (data not shown) verifies that the sequences from the different species are, in fact,orthologous. The tree and branches are derived directly from Eizirik et al.(2001).











