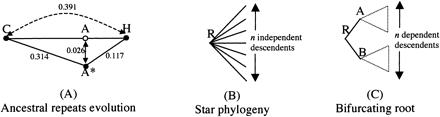
(A) Estimates of the expected number of substitutions per site between a repeat consensus C, it human descendent H, and the reconstructed ancestor A*, based on a Kimura 2-parameter model and averaged over all human ancestral repeats of the region considered. The true ancestor A cannot be observed, but a distance of 0.026 substitutions per site between it and A* is estimated from the three other distances. (B) Star phylogeny with n independent descendants. (C) A tree with bifurcating root. Irrevocable information loss occurs between R and its descendants A and B.











