Average observed/expected (O/E) values for the three models explaining codon usage in slow growers and fast growers
|
|
Fast growers |
Slow growers |
||||
|---|---|---|---|---|---|---|
|
|
dif |
All |
dif |
All |
||
| Frequency | 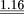
|
1.04 | 
|

|
||
| Perfect match | 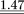
|

|
1.03 | 0.99 | ||
| Stability
|

|
0.93
|
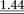
|
0.94
|
||
-
Note that because expected values are generally different in the different models, O/E values between different models cannot be compared in a straightforward manner. “All” corresponds to the fit when the most frequent codon per amino acid is taken from the codon usage of all genes. “Dif” corresponds to the fit when the most frequent codon per amino acid is the one increasing the most in frequency in ribosomal proteins compared to the rest of the genome. Wilcoxon tests (Ha: O/E > 1), underlined values (P < 0.05), double underlined values (P < 0.001).











