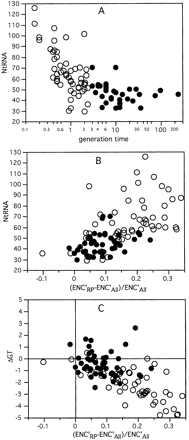
(A) Number of tRNA genes per genome as a function of the bacterial optimal generation time. (B) Number of tRNA genes per genome as a function of codon usage bias given as the difference in ENC′ between the set of genes coding for ribosomal proteins (ENC′RP) and the whole genome (ENC′All). (C) Enrichment in G and T in the last codon position of twofold-degenerated amino acids in ribosomal proteins compared to the average genome (ΔGT) as a function of codon usage bias. ○, fast growers; •, slow growers (generation time >2.5 h).











