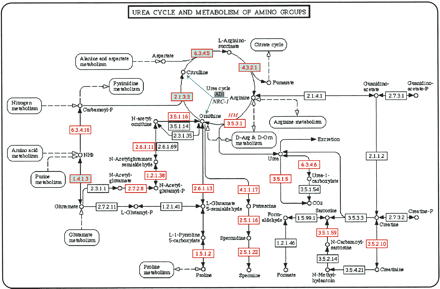
Comparative metabolic reconstruction in H. marismortui and Halobacterium sp. NRC-1. An example of metabolic reconstruction using the Web intermediary software and KEGG: arginine metabolism and the urea cycle in H. marismortui compared with Halobacterium sp. NRC-1. Enzymes identified in Halobacterium sp. NRC-1 are shaded in green, and those identified in H. marismortui are shown with a red border and red font. The ADI pathway of arginine fermentation in Halobacterium sp. NRC-1 is indicated with solid blue arrows. Each enzyme identified in H. marismortui is hyperlinked to its corresponding entry in the SBEAMS annotation database. This view of KEGG metabolic pathways for the two halophiles is available on http://halo.systemsbiology.net.











