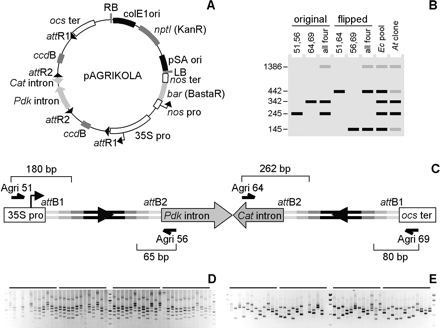
Construction of hpRNA expression plasmids. (A) Map of the pAGRIKOLA destination vector. (B) Schematic representation of the multiplex analysis of the expression plasmids. Numbers on the left show how many base pairs were added to the primary GSTs upon amplification with the Agri primers indicated at the top of each lane. The largest band corresponds to the amplification of both GST cassettes with the external Agri51 and Agri69 primers. Profiles in lanes 7 and 8 correspond to results in panels D and E, respectively, and are generated via amplification with all four Agri primers. The profile in lane 8 only shows results obtained with a plasmid in which the spacer intron is initially in its original orientation, as pictured in C. (C) Structure of recombined hairpin cassette with the inverted GST repeats. GST elements are depicted as in Figure 3. Bracket numbers indicate the length of the fragment delimited by the 5′-end of each Agri primer and the 3′-end of the closest attB site, on either strand. Configuration with the flipped spacer intron is not shown. (D) Multiplex PCR validation of double LR clonase reaction products. Each lane shows the amplification results for one E. coli pool after liquid selection (see Supplemental Table S2). (E) Multiplex PCR verification of A. tumefaciens transformants. For each initial pool, four individual Agrobacterium colonies were picked. Most tetrads contain both types of colonies, with the expression plasmid carrying the intron spacer initially either in the original or flipped orientation (see Supplemental Table S2).











