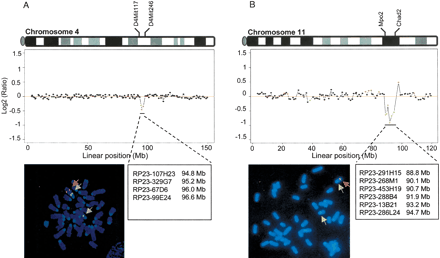
Detection of regional single-copy deletions. (A) Comparative genomic hybridization of normal AB2.2 ES cell DNA vs. DNA from 7A9 ES cells, harboring a regional single-copy deletion between D4Mit117 (94.6 Mb) and D4Mit246 (96.6 Mb) on chromosome 4. (B) Comparative genomic hybridization of normal AB2.2 ES cell DNA vs. DNA from 3D5 ES cells, harboring a 6.9-Mb single-copy deletion from Mpo2 (88.5 Mb) to Chad2 (95.4 Mb) on chromosome 11. The BAC clones spanning the deletion regions (green dots) are denoted. Both deletions were confirmed by FISH analysis. On prometaphase chromosomes from 7A9 and 3D5 ES cells, only one hybridization signal was observed, with probes corresponding to the deleted region (red signals, indicated by red arrow), whereas two hybridization signals, derived from both alleles, were detected with probes located outside of the deletion regions (green signals, indicated by green arrows).











