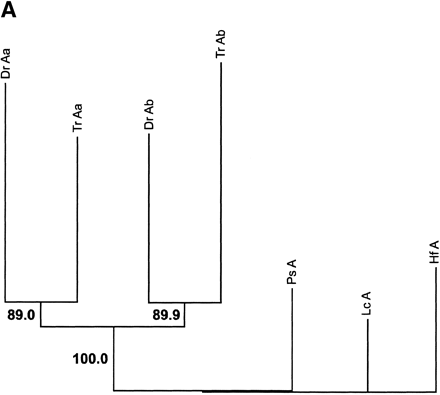
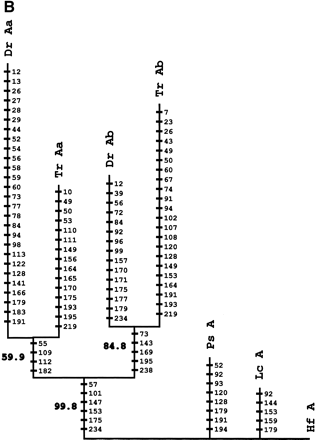
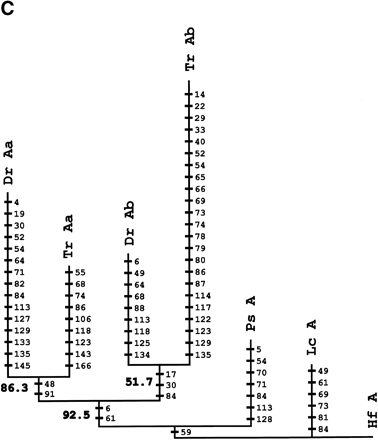
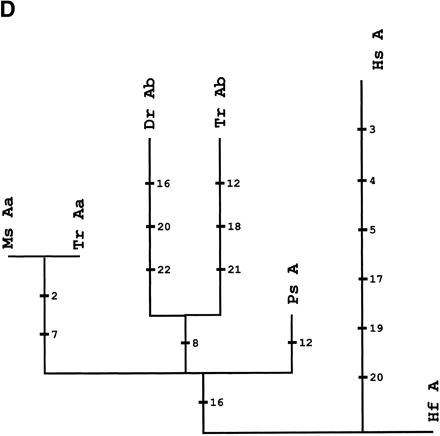
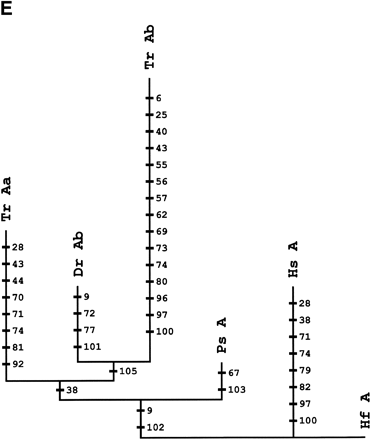
Phylogenetic analysis andcharacter reconstructions of Hox coding sequences. (A) Neighbor-joining tree of concatenated HoxA13 and HoxA11 exon 1 coding sequences. Bootstrap support (1000 replications) for the euteleost nodes are shown. (Hf) Heterodontus francisci; (Lc) Latimeria chalumnae; (Ps) Polypterus senegalus; (TrAb) Takifugu rubripes HoxAβ; (DrAb) Danio rerio HoxAβ; (TrAa) T. rubripes HoxAα; (DrAa) D. rerio HoxAα. (B-E) Character reconstructions of exon 1 coding sequences under constraint trees for HoxA13, HoxA11, HoxA10, and HoxA2, respectively. Taxa abbreviations are as in A above, including (Hs) Homo sapiens. The indicated substitutions are only those that map unambiguously to the branches of the trees. The numbers of analyzedamino acidresidues andassumedunambiguous changes for each gene are (B) HoxA13, 244 amino acids and 196 steps; (C) HoxA11, 192 amino acids and 151 steps; (D) HoxA10, 24 amino acids and 35 steps; (E) HoxA2, 105 amino acids and 59 steps.











