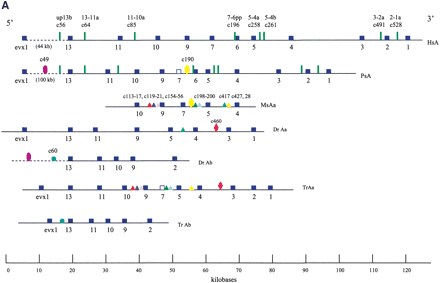
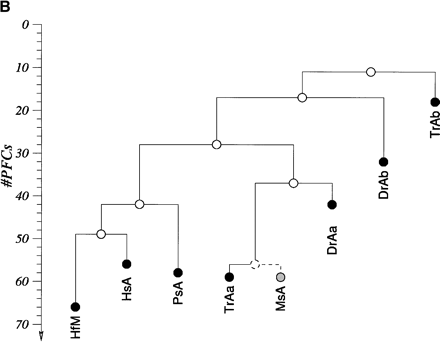
HoxA clusters and PFCs in human, shark, bichir, andeuteleosts. (Hs) Homo sapiens; (Hf) Heterodontus francisci; (Ps) Polypterus senegalus; (MsAa) Morone saxatilis HoxAα; (DrAa) Danio rerio HoxAα; (DrAb) D. rerio HoxAβ; (TrAa) Takifugu rubripes HoxAα; (TrAb) T. rubripes HoxAβ. (A) Hox genes are indicatedby blue rectangles. PFCs sharedexclusively between human andbichir are indicatedby green bars. PFCs sharedexclusively between bichir andeuteleosts are indicatedby coloredellipses. PFCs sharedbetween the HoxAα clusters of striped bass, zebrafish, and/or pufferfish are indicated by coloredtriangles. The PFC sharedbetween only zebrafish andpufferfish HoxAα clusters is indicated by a red diamond. The PFC shared between zebrafish andpufferfish HoxAβ clusters is indicated by an aqua blue circle. See text for description. (B) The co-occurrences of the PFCs in different clusters (Supplemental Table 1) can be representedas a tree. The height of an internal node is the average number of PFCs shared by two clusters in the two different subtrees. The position of the tips gives the total number of PFCs in the segment of the HoxA cluster that spans from Evx1 to HoxA1. The nonduplicated HoxA regions of human, shark, andbichir form one group. The secondsignificant group consists of the HoxAα sequences of the euteleosts. The position of the incomplete stripedbass (Morone saxatilis) sequence is estimatedby assuming that in a complete sequence we wouldhave foundroughly the same number of PFCs as in pufferfish (T. rubripes). The HoxAβ clusters are much further divergedanddo not appear to group together because euteleost-specific PFCs in the HoxAβ cluster are very rare (Supplemental Table 1). The smaller rate of PFC loss in the zebrafish HoxAβ cluster is the dominating effect.











