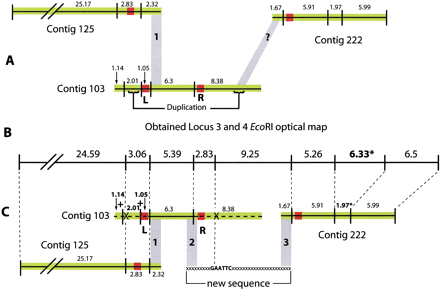
The melding of contigs 125, 103 and 222 after the resolution of contig 103 misassembly by optical mapping, see also Locus 3 and 4 in Table 2. (A) The three contigs 125, 103 and 222 are shown in the order suggested by sequence scaffold data. Contigs are represented by black horizontal lines. In silico EcoRI sites are represented by vertical lines and between them the predicted in silico EcoRI fragment sizes in kb. Green areas indicate regions of high sequence quality (10–22 -fold coverage), red areas of poor sequence coverage (2–4 fold) and/or poor quality sequence data. Grey block 1 indicates a region of high sequence identity between the 3′ end of contig 125 and a region lying 4.5 kb from the 5′ end of contig 103. The angled grey block indicated by a question mark (?) shows a region of high sequence identity between the 5′ end of contig 222 and a region towards the 3′ end of contig 103. This match later proved to be spurious. The 3′ region of contig 103 is duplicated internally towards its 5′ end. (B) The EcoRI optical map covering part of the region for loci 3 and 4. The sizes in this panel are optical map fragment sizes in kb. (C) The solution to the resolution of this region. EcoRI sites considered equivalent on the optical and in silico maps are joined by fine dotted lines. Optical mapping indicated that poor quality region L had resulted in a misassembly at the 5′ end of contig 103. This was concluded from: 1) the presence in a region of high quality sequence of two additional EcoRI sites (indicated by + signs); and 2) the absence of a restriction site (indicated by X). These were in conflict with the optical map data. Removal of the sequences 5′ of and inclusive of, region L (indicated by a dotted horizontal line) permitted the melding of contig 125 and 103 at grey region 1. The absence of an EcoRI site 3′ of poor quality region R (represented by X) suggested that region R may have given rise to a misassembly of the 3′ end of contig 103. Removal of this region (also indicated by a dotted line) followed by resequencing in this area resulted in the acquisition of a missing EcoRI site and permitted the assembly of the new sequence and joining of contigs 103 and 222 at grey regions 2 and 3. Note that contig 222 has a 1.97 kb EcoRI fragment; however, optical mapping suggests this fragment should be 6.33 kb (these fragments are marked *). To resolve this size discrepancy additional sequencing of this region is currently under way.











