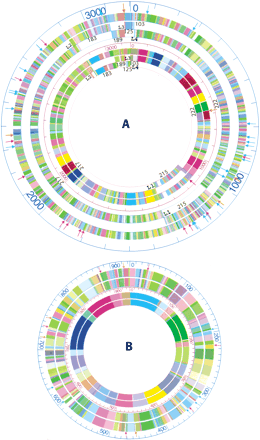
The alignments of the HindIII and EcoRI consensus optical maps with corresponding in silico maps. (A) The outer three rings show the alignment of the in silico EcoRI optical map (outermost ring) with the chromosome I in silico maps (7 total; numbered), while the inner three circles show the alignment of the HindIII optical map (the outer one) with the chromosome I in silico HindIII maps. The numbers indicate the sequence contig number. L1, L2, L3, and L4 indicate the four loci for adjacent sequence contigs, and detailed map alignments are shown in Tables 2 and 3. Orange arrows denote false cuts, red arrows denote missing cuts, and light blue arrows denote missing fragments (missing fragments less than 1 kb, or one of the two fragments for missing cuts and false cuts were not shown). (B) The outer two rings show the alignment of the EcoRI in silico map with the corresponding optical map (the outermost ring) of chromosome II, while the inner two rings show the alignment of the HindIII in silico map with the HindIII optical map (outer). Orange arrows denote false cuts, red arrows denote missing cuts, and light blue arrows denote missing fragments (missing fragments less than 1 kb, or one of the two fragments less than 1 kb for missing cuts and false cuts are not shown).











