Comparison of Strategies A and B for Fine Mapping the Location of the Disease Mutation in 100 Data Sets Simulated Based on the Cystic Fibrosis Data
|
LD mapping algorithm |
Strategy |
Mean(pos)a |
Std(pos)b |
Mean 95% PI widthc |
RMSEd |
Overlape |
|---|---|---|---|---|---|---|
| BLADE | A | 0.8600 | 0.0145 | 0.1461 | 0.0262 | 99% |
| B | 0.8647 | 0.0315 | 0.1402 | 0.0349 | 96% | |
| DHSMap | A | 0.8868 | 0.0428 | 0.2141 | 0.0432 | 96% |
|
|
B
|
0.9072
|
0.0702
|
0.1620
|
0.0749
|
66%
|
-
↵a The average of the 100 location estimates in the 100 simulations (true location = 0.88 cM).
-
↵b The sample standard deviation of the 100 location estimates.
-
↵c The average width of the 95% probability intervals (PIs) in the 100 simulations.
-
↵d Root mean square error (RMSE): square root of the average squared differences between the estimated and the true location, i.e., RMSE =
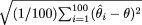 , where θ is the true location and
, where θ is the true location and 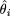 is the location estimate based on the ith simulated data set.
is the location estimate based on the ith simulated data set.
-
↵e The percentage of times (in the 100 simulations) that the 95% PI overlapped with the target region.











