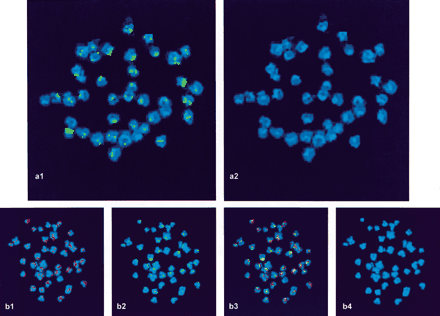
Chromosomal localization of Zebulon in the genome of T. nigroviridis by FISH. Weak, scattered spots have been removed by electronic thresholding in order to retain only major regions of accumulation. The genomic areas in which Zebulon preferentially localizes (a1) mostly correspond to heterochromatic, DAPI-positive regions as shown in this over-denaturated metaphase (a2). Double FISH between Zebulon (DIG-labeled pG16H23, b1) and Rex3 (biotin-labeled, b2), a non-LTR retrotransposon abundant in the genome of T. nigroviridis (C. Fischer, L. Bouneau, and C. Ozouf-Costaz, unpubl.) shows superimposed signals (b3) corresponding to common regions of accumulation in DAPI-positive regions (b4).











