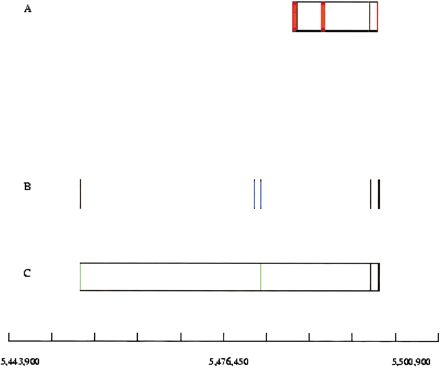
Ecores correcting a gene model. The scale refers to the position on the chromosome arm 3L of the genome of Anopheles. (A) Ensembl gene predictions (release 6.1a) on the 5′-3′ strand. The genes are represented by boxes, with vertical lines separating exons (red) and introns (white). (B) Ecores (blue). (C) A cDNA sequence is in green, with potential introns in white. Only one cDNA, matching with unannotated ecores, is represented here. This cDNA (corresponding to the assembly of entries BX062803 and BX062804) matches two of the three orphan ecores. It is homologous throughout to a Drosophila tetraspanin family member. The version 6.1a of the annotation apparently fused the two last exons of the gene with two putative exons, originating from a transposable element. The large sizes of the two first introns may induce such erroneous model constructions. In release 10.2.1, the region is entireley unannotated.











