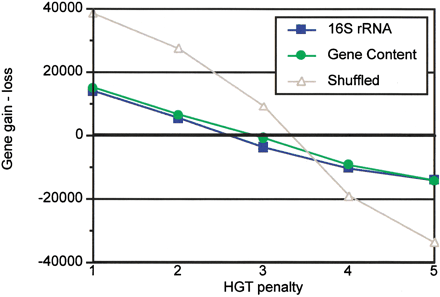
The difference between gene gain and loss at various HGT penalties. The results for 16S rRNA, gene content, and shuffled trees are shown; represented as in Figure 1. The concordance of the two genuine (16S rRNA and gene content) trees is evident, contrasted to the shuffled tree. Genome stability (signified by zero difference, bold horizontal line) is achieved with HGT penalty values between 2 and 3.











