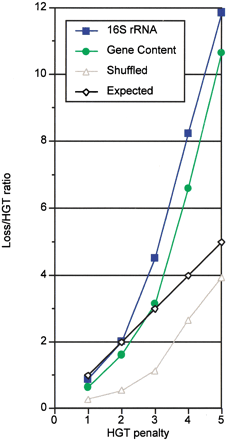
Comparison of observed and expected ratio of gene loss over HGT at various HGT penalties. The expected ratio implied by the model is shown in black (diamonds), and observed results are represented in blue (squares) for the 16S rRNA tree, green (circles) for the gene content tree, and gray (triangles) for the shuffled tree. The genuine trees are in agreement with the model (intersecting the expected linear curve) at HGT penalty values between 2 and 3. The shuffled tree overpredicts HGT at any tested HGT penalty value.











