Functional Annotation of RIKEN Mouse cDNA Clones Using GNF Expression Atlas
Annotation of novel cDNAs relies on multiple lines of evidence, including gene structure, predicted ORFs, and predicted protein motifs. Insight into function also relies on expression profiling, through querying EST databases of direct experimentation.
High-throughput gene expression profiling, by use of spotted arrays or oligo chip methods, has become an important tool for investigating transcriptional activity. Among cDNAs predicted to encode related proteins, different expression patterns can suggest functional specialization. Additionally, common expression patterns can serve to group otherwise unrelated cDNAs. Expression profiling may be particularly useful for difficult to annotate clones, for which other lines of evidence are not available.
GNF has generated and analyzed gene expression previously from a set of 45 murine samples across a diverse list of normal tissues, organs, and cell lines probed with the Affymetrix MG-U74A chip. These data have been published (Su et al. 2002) and are available at the GNF's free and publicly accessible Web site (Gene Expression Atlas, http://expression.gnf.org), which integrates data visualization and curation of current gene annotations.
We identified the relationship between the RIKEN collection and MG-U74A/B/C targets by SIM4-polished BLAST alignments between 60,770 RIKEN clone sequences and 36,701 target sequences arrayed on the U74A/B/C chips. The results of this analysis are summarized in Table 1. There are 25,831 RIKEN clones represented on this array set. Of these, 2994 have lower quality similarity hits at the DNA or protein level (i.e., are not automatically or easily annotated by homology).
The Counts of RIKEN Clones Represented on MG-U74 A, B, and C Chips
Ten additional tissues were profiled using the Affymetrix MG-U74Bv2 and MG-U74Cv2 chips to allow for inferred functional annotation in the absence of other evidence. These tissues were chosen to give the greatest representation of gene expression by extrapolation from the results obtained with U74A. Five of the tissues are also in common with those used to probe the RIKEN microarray. The combined resources should provide an extremely comprehensive characterization of the expression data.
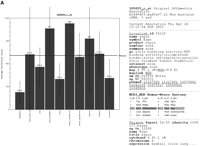
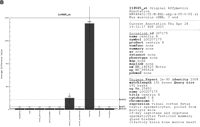
Figure 1 shows just two arbitrarily taken cases in which tentative annotation may be inferred from the expression profile.
(A) RIKEN clone 0610007F01 is interrogated by the probeset 165655_r_at. (B) RIKEN clones 6430560A18 and G430087M20 are associated with the probe set 114585_at with testis-specific pattern.
Footnotes
-
Article and publication are at http://www.genome.org/cgi/doi/10.1101/gr.1457103.
-
↵1 Corresponding author. E-MAIL batalov{at}gnf.org; FAX (858) 812-1570.
- Cold Spring Harbor Laboratory Press