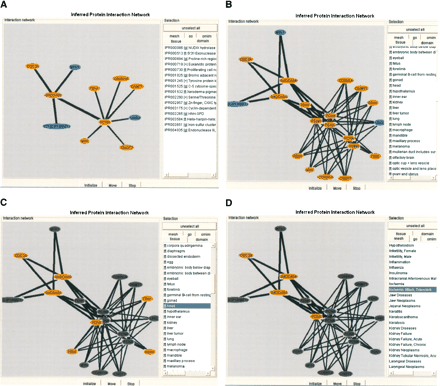
(A) MEDLINE abstract-derived PPI network for curated PPI of 11 proteins: CDC2A, CDKN1A, CNOT7, FEN1, GADD45A, PCNA, MTK1, P21(CIP1/WAF1), UNG2, MYH, and DNMT1 (see also http://facts.gsc.riken.go.jp/viewer/InteractionViewer.php?CloneID=2810049I05 for protein interaction viewer of clone ID 2810049I05). Detailed information on data source accessions and inferred functions (OMIM Morbidmap, disease MeSH, tissue distribution, gene ontology assignment, and InterPro domains) appears after clicking on the circles. Orange-colored circles symbolize mouse proteins. Blue circles show potential interacting proteins of human origin. (B) The PPI network shown in A is derived from 64 interaction pairs inferred from 14 FANTOM2 and 5 GenPept sequences. The clone-based display of inferred PPI facilitates the visualization of context information. (C) Selection of head shows CDC2A—GADD45A—PCNA—FEN1 PPI associated with transcripts that are expressed in the head. Symbols preceding the tissue names, (f) FANTOM2 READ database-derived EST tissue information; (g) dbEST-derived EST tissue information. (l) FANTOM2 cDNA library source-derived tissue information. (D) Additional selection of disease MeSH Ischemic attack, transient removes FEN1 from the PPI network.











