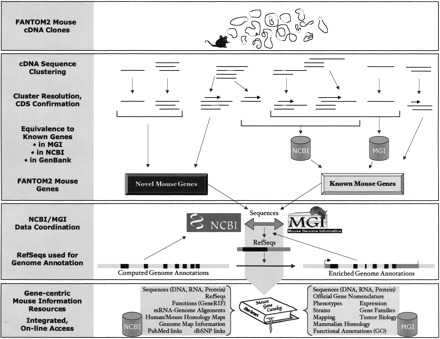
The flow from FANTOM2 mouse cDNA clones to genes, to their integration with NCBI LocusLink and MGI. Mouse cDNA clones isolated (closed circles in the first panel) and sequenced (horizontal lines in the second panel) by the RIKEN group are clustered computationally (top clusters in the second panel). Computed clusters are then resolved into gene-specific groups by human inspection (bottom clusters in the second panel). Dotted lines represent transcript variation. Computed clusters can group sequences from different genes, such as paralogs and read-through transcripts (third and fourth computed clusters from left, respectively), and other distinct gene sequences that share some region of overlap requiring manual resolution. CDS regions for protein coding genes are indicated (horizontal arrows over clusters). Equivalence of FANTOM2 sequences with known mouse genes in NCBI LocusLink and MGI is detected by incorporation of known sequences in the FANTOM2 clusters or by BLAST (data not shown). LocusLink and MGI contain overlapping but distinct sequence data sets. Some characterized mouse sequences not present in LocusLink or MGI can have sequence identity to FANTOM2 sequences (far right cluster). Remaining FANTOM2 genes are considered novel. The curation of sequences for novel and known mouse genes is coordinated between LocusLink and MGI, and LocusLink establishes RefSeqs (third panel). Genome centers feed predicted gene models to NCBI, but rely on transcript-based evidence in the form of RefSeqs to improve genome annotations. Gene models with enriched annotations link back to gene records in LocusLink and MGI on the basis of integrated sequence accessions. Through data coordination, LocusLink and MGI establish a catalog of mouse genes with accurate sequence associations and integrated biological information.











