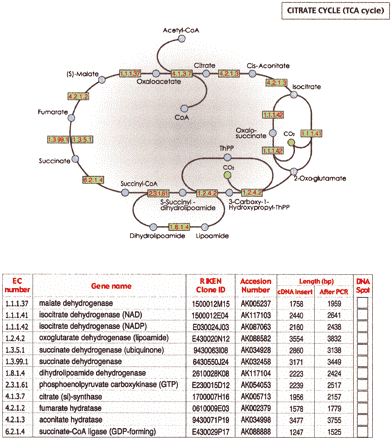
A DNA sheet is attached as a sample for field testing. Twelve cDNA plasmids of the RIKEN Mouse Genome Encyclopedia annotated as genes involved in TCA cycles (Bono et al. 2003) are attached in the right column of the annotation table. EC numbers and gene names are used according to the KEGG database (Kanehisa et al. 2002). The cDNAs were cloned into the pFLC vector (Carninci et al. 2001), which is identical to the pBluescript except for the modified cloningsites. A total of 100 ngof plasmids were spotted on water-soluble MDP60 paper (Mishima Co., Ltd.), and dried. Positions of attached plasmids are shown in red. TCA cycle and EC number of genes are shown at top. To amplify cDNA inserts, cut out the 2 mm × 2 mm area spotted with DNA, place it into a PCR tube, add 50 μL of PCR solution, centrifuge the resulting solution, and then initiate PCR cycle. PCR solution contains 1.5 U of KOD Plus DNA Polymerase (TOYOBO Co., Ltd.), and 1× KOD Plus PCR buffer together with 0.2 μM of PCR primers (-21M13; 5′-TGTAAAACGACGGCCAGT-3′, 1233-Rv; 5′-AGCGGATAACAATTTCACACAGGA-3′), 0.2 mM each of dATP, dGTP, dCTP and dTTP, and 5.3 mM MgCl2. After centrifuging the resulting solution, the PCR cycle is initiated. PCR cycles comprise 2 min at 94°C; 29 cycles of denaturing(94°C, 1 min), annealing(60°C, 1 min) and extension (68°C, 75 sec), and 15 min at 74°C. Alternatively, an aliquot of soluble-paper solution can be subjected to E. coli transformation. This sample DNA sheet is attached for large-scale field testing of the DNA Book, to test how well the DNA sheet is delivered to readers. We would gratefully appreciate readers attempts to extract genes printed on this sheet. Feedback can be input on our Web page, http://genome.gsc.riken.go.jp/DNA-Book/. Results from readers will be reported as a separate paper in the near future. Patent for DNA Book pending.
J. Kawai and Y. Hayashizaki











