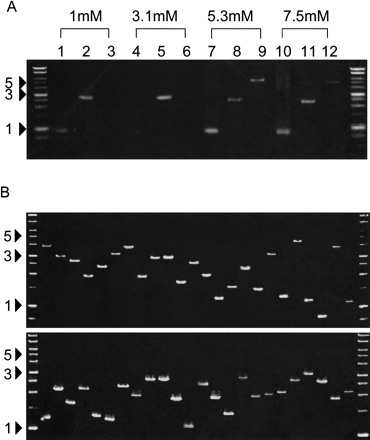
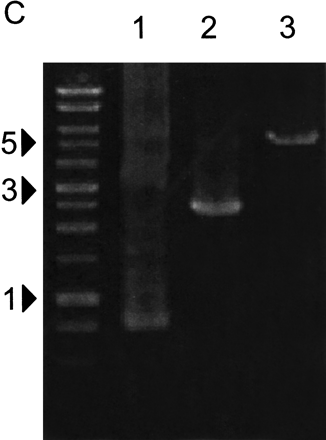
Extraction of DNA from DNA sheets by PCR. (A) PCR conditions for DNA extraction. Fifty nanograms of three RIKEN mouse cDNAs (722 bp, lanes 1,4,7,10; 2418 bp, lanes 2,5,8,11; 5438 bp, lanes 3,6,9,12) in 0.5 μL plasmid solution were spotted on water-soluble 60MDP paper (Mishima Co., Ltd.). After dryingin air for 30 min, insert DNA was amplified from paper pieces (4 mm × 4 mm) using PCR with vector-specific primer in the presence of various concentrations of Mg2+. DNA marker sizes are shown in kilobases. (B) Various DNA inserts can be recovered from DNA sheets. Random selections of 93 plasmids of RIKEN mouse cDNA clones, ranging in size from 732–4896 bp, were extracted from DNA sheets. In two independent experiments, almost all cDNA inserts (95% and 100% of 93 clones) were amplified successfully. Representative results for 24 samples are shown. Marker sizes are shown in kilobases. (C) DNA sheets can tolerate rubbingat 37°C in 70% humidity overnight. DNA sheets spotted with ∼50 ngof cDNA plasmids were kept in a humidified incubator or were inserted into an issue of Genome Research, and shaken overnight at 180 rpm under conditions of 37°C and 70% humidity. DNA inserts of cDNA were extracted from DNA sheets using PCR, as described in the Methods. Lanes 1, 2, and 3 represent 722, 2418, and 5438 bp cDNA inserts, respectively.











