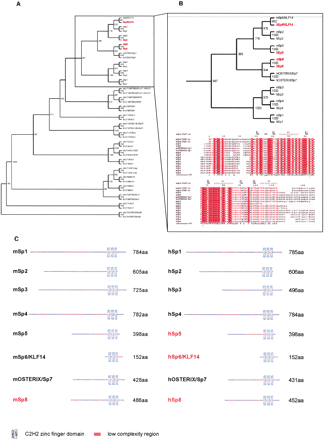
(A) Unrootedphylogeny among the Sp/Krüppel-like factors. The entire mouse and human protein sequences of the Sp/Krüppel-like factors (Table 4) were aligned and subjected to Neighbor Joining with 1000 bootstrap analysis. (B) Magnification of the Sp branch of the phylogeny tree and alignment of the zinc finger region of the Sp proteins. The secondary structure of the six fingers is shown above the alignment; the domain consensus is shown below the alignment. (C) Mouse and human Sp protein domain architecture. Those highlighted in red are the newly described proteins.











