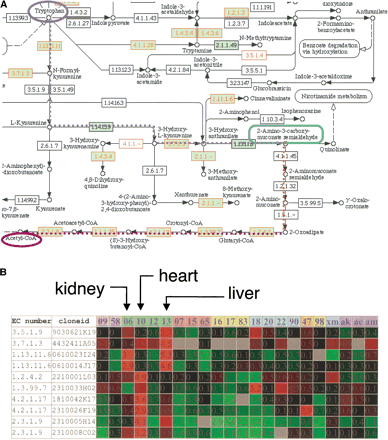
Reconstruction of mouse tryptophan metabolism pathway in silico. (A) An example of an automatically reconstructed pathway using the KEGG Web interface by specifying multiple EC numbers in the query. The EC number in the green box was found in the human metabolome set, and the EC number drawn in red was covered by the RTPS. There were missing enzymes on the pathway that were assigned computationally. These enzymes are described in Table 2. Four of six missing clones were finally identified after intensive human curation. (B) Gene expression of mRNAs coding enzymes in the tryptophan metabolism pathway by microarray. Genes are sorted by order of the cascade of tryptophan degradation to Acetyl-CoA, not by clustering by similar expression profiles. It is apparent that the genes in the upper stream of this cascade are highly expressed in liver and kidney, and those downstream of the cascade are highly expressed in heart. 09, spleen; 58, thymus; 06, kidney; 10, heart; 12, lung; 13, liver; 07, brain; 15, cerebellum; 65, cerebellum neonate 10 day; 16, placenta; 17, testis; 83, uterus; 18, pancreas; 20, small intestine; 22, stomach; 90, colon; 47, skin neonate 10 day; 98, bone; xm, muscle; ak, adipose dorsal kidney; ae, adipose epididymal; am, adipose mesenteric.











