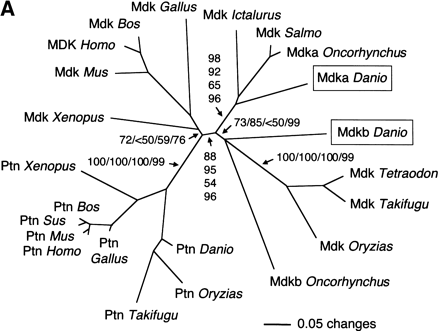
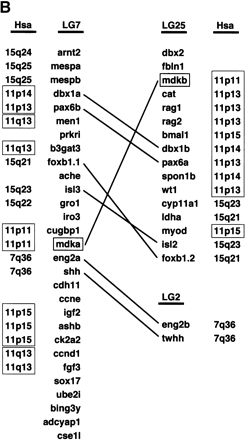
(A) Phylogenetic analysis of midkine and pleiotrophin proteins. The amino acid tree (neighbor-joining) is unrooted. Bootstrap values using neighbor-joining (first and second values) and maximum parsimony analyses (third values), as well as reliability values for maximum likelihood quartet puzzling (fourth values) are given. Neighbor-joining was performed either considering all sites (first values) or only the unsaturated fraction of sites (second values) using AsaturA (Van de Peer et al. 2002). Branches with <50% support have been collapsed. Accession nos. as follows: PTN Homo sapiens CAA37121; Ptn Mus musculus AAH02064; Ptn Bos taurus A37780; Ptn Sus scrofa BAA13972; Ptn Gallus gallus P32760; Ptn Xenopus laevis BAA07659; Ptn Oryzias latipes AU170438 (EST); Ptn Danio rerio AL925602 (EST); Ptn Takifugu rubripes scaffold S001533 (http://fugu.hgmp.mrc.ac.uk); Mdk Bos taurus BAA98139; the origin of other Mdk sequences is given in the legend to Fig. 1A,Fig. 1B. (B) Zebrafish mdka and mdkb are located on two different linkage groups (LG) presenting synteny with human chromosome Hsa11. Duplicate genes on different linkage groups are shown. Radiation hybrid mapping was done using the LN54 mouse-zebrafish panel (Hukriede et al. 2001). Only putative orthologies on Hsa11, Hsa15, and Hsa7 are shown. Putative orthologies were determined by using information available from the ZFIN server (http://zfin.org) and by searching the draft of the human genome using the zebrafish protein sequences as TBLASTN queries (http://www.ncbi.nlm.nih.gov/BLAST).











