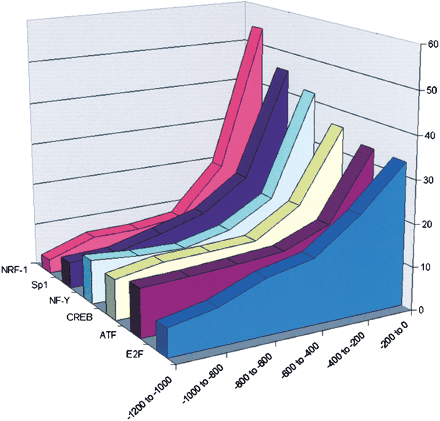
Distribution of locations of TF putative binding sites found in 568 cell-cycle-regulated promoters. Promoters were divided into six intervals, 200 bp each. For each of the PWMs listed in Table 3, the number of times its computationally identified binding sites appeared in each interval was counted (after accounting for the actual number of base pairs scanned in each interval; this number changes as the masked sequences are not uniformly distributed among the six intervals). Locations of NRF-1, CREB, NF-Y, Sp1, ATF, and E2F binding sites tend to concentrate in the vicinity of the TSSs (χ2 test,p < 0.01).











