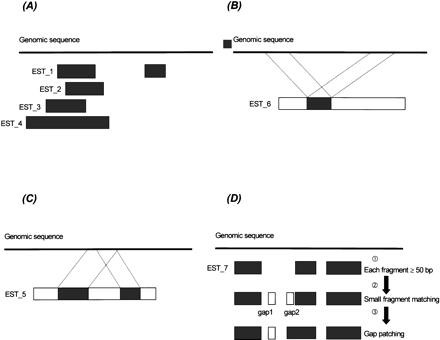
Four possible scenarios of ESTs matched to a genomic sequence by CRASA. (A) Multiple ESTs match to the same region of genomic sequence. (B) Several internal segments of an EST match to the same region of a genomic sequence. (C) One segment within an EST matches to several regions of a genomic sequence. (D) The potential coding region of a genomic sequence matches colinearly to an EST in split segments and the processes of small fragments matching and gap patching. Step 1: EST_7 has three segments matched to the genomic sequence, which are at least 50 bp in length. Step 2: Small fragment matching: the small matched fragments (10–49 bp) are shown (open box). Step 3: Gap patching with the genomic sequence: While Gap 1 stays independently as a match, Gap 2, corrected by the gap-patching rule, is patched contiguously to the matched fragment downstream.











