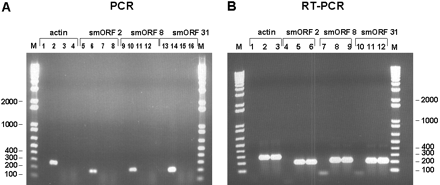
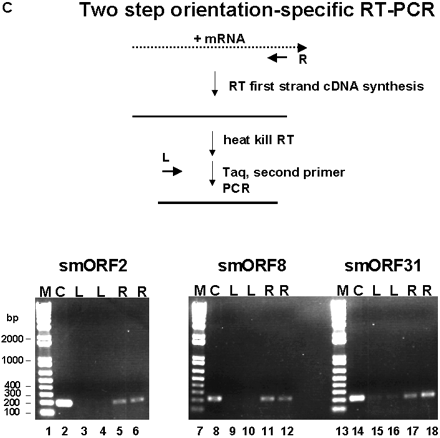
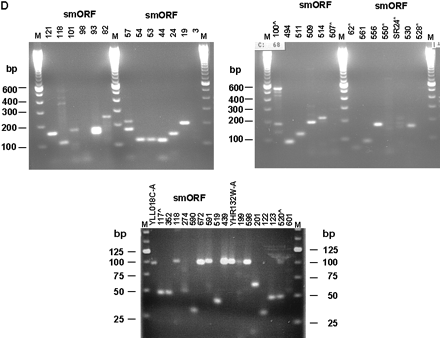
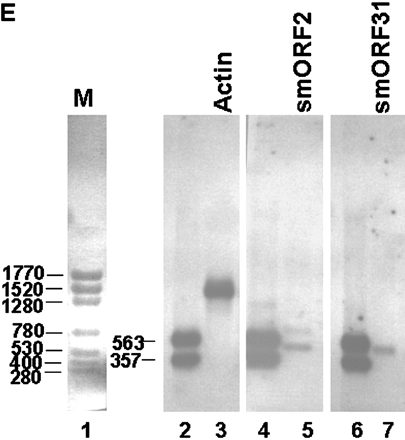
Experimental validation of the S. cerevisiae smORFs. (A) Primers specific for the yeast ACT1 gene as well as the three smORFs were used for PCR amplification using no template (lanes 1,5,9,13), 50 ng genomic DNA (lanes 2,6,10,14), 500 ng total RNA from cells grown in rich media (lanes3,7,11,15), and 500 ng total RNA from cells grown in minimal media (lanes4,8,12,16) .(B) Primers specific for the yeast ACT1 gene as well as the three smORFs were used for RT-PCR amplification using no template (lanes1,4,7,10), 500 ng total RNA from cells grown in rich media (lanes2,5,8,11), and 500 ng total RNA from cells grown in minimal media (lanes3,6,9,12). PCR and RT-PCR products were fractionated on a 1% agarose gel with DNA size markers and visualized after ethidium bromide staining. Sizes of DNA fragments in bp are indicated. (C) Two-step orientation-specific RT-PCR. Primers whose sequence is complementary to the predicted mRNAs of smORF2, 8, and 31 were used for first-strand cDNA synthesis. After heat inactivation of the reverse transcriptase, PCR amplification was carried out with both smORF-specific primers (lanes5,6,11,12,17,18). As control, the experiment was repeated using primers with the same sequence as the mRNA for first-strand cDNA synthesis (lanes3,4,9,10,15,16). (D) Examples of RT-PCR results with various smORFs indicated on top. RT-PCR detection of transcripts from the annotated smORFs YLL018C-A and YHR132W-A are shown. smORFs for which RT-PCR reactions resulted in no products are indicated (*) as are those for which the product is not of the expected size (∧). These smORFs were not included in Table 1. Sizes of DNA fragments in bp are indicated. (E) Transcript size determination by Northern analysis of the ACT1 (lane 3), smORF2 (lane 5), and smORF31 (lane7). Unlabeled RNA markers stained with methylene blue (lane1) and labeled RNA markers (lanes2,4,6) were fractionated together with yeast poly (A)+ RNA. Sizes are shown in nucleotides.











