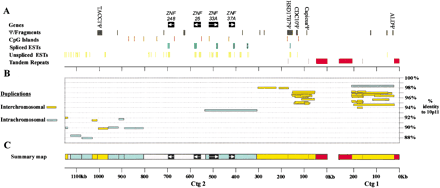
Overview of 10p11 sequence data. (A) Principle features of sequence: The position of known genes (Table 1) and pseudogene fragments (http://www.ncl.ac.uk/ihg/10p11.htm), CpG islands, spliced and unspliced ESTs (Table 1), and tandem repeats (Table 3). The orientation of gene transcription is indicated with white arrows. (B). Paralogs of the 10p11 sequence within the human draft. These have been divided into interchromosomal (yellow) and intrachromosomal (blue). The position of each independent BLAST hit greater than 5 kb in length within the EMBL-NR and EMBL-HTG databases are represented by a horizontal line. Both the extent of the BLAST hit within the 10p11 sequence (x-axis) and the % identity to the 10p11 sequence (y-axis) is indicated (see Methods). Deletions and insertions within individual sequence matches are not shown. (C) Summary map showing genes, satellite repeats, and duplications for reference with later Figures. Because satellite repeats anchor this map, the physical scale of each contig is indicated in 100 kb intervals moving away from the centromere towards 10pter (right toleft). For details of sequence analysis see Methods. The sequence composition of the ∼27–600 kb gap between contigs 1 and 2 is unknown.











