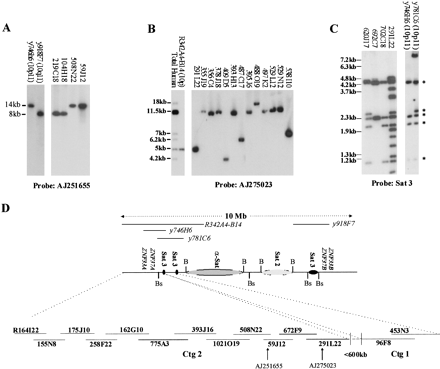
Construction of 10p11 contigs. (A–C) Southern hybridization analyses used to assess chromosomal/regional origin of BACs. The map position/sequence content of reference YACs (y746H6, y918F7, and y781C6) and a somatic cell hybrid (R342A4-B14) is given in parenthesis, and fragment sizes are given in kilobase pairs (kb). All DNAs were digested with EcoR1. The 10p11 probes are defined by their accession numbers and the satellite 3 probe used was pHS5 (Cooke and Hindley 1979). The regional localization of all BACs sequenced was confirmed using these resources in combination with FISH analyses (data not shown). (D) BAC tiling path in relation to known pericentromeric satellite organization. The approximate position ofBssHII (Bs) and BamH1 (B) sites which define the satellite arrays (Jackson et al. 1993), and the duplicated ZNF genes which flank the centromere (Tunnacliffe et al. 1993), are shown. The content of YACs and somatic cell hybrid used in panels (A–C) is also indicated above the satellite map. All BACs in both contigs are from the RPCI11 library, with the exception of R164I22, which was obtained from Research Genetics (see methods). The orientation of contig 1 and the estimate of the gap size between the contigs is based on sequence identity, number, size, and known position of satellite arrays (see text). The full depth contig from which tiling path 2 was chosen can be viewed athttp://www.ncl.ac.uk/ihg/10p11.htm.











