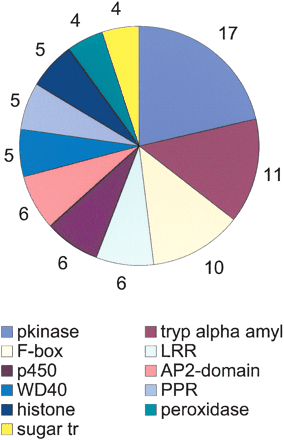
The most common domains in tissue-enriched SASs. The SASs were inspected for tissue specificity using the library of origin of their EST components. An SAS was considered tissue-enriched when it contained at least three ESTs found exclusively in a given library. Of the 43,141 SASs, 1234 were tissue-enriched and were inspected for the presence of conserved protein domains. The SASs were translated using the ESTScan algorithm. Domain occurrence was revealed by querying the Pfam database with the corresponding amino acid sequences using the default settings of Pfam 7.0 [“global and local alignments merged” and “Pfam gathering threshold (GA)”]. The most represented domains among the tissue-enriched SASs along with the number of SASs for each of then are shown.











