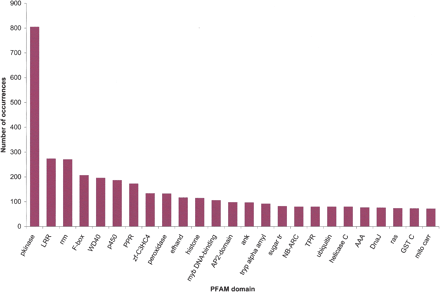
The number of occurrences for the 25 most common Pfam domains in SAS proteins. The 43,141 SASs were translated using the ESTScan algorithm, and the resulting 40,821 amino acid sequences were then entered as queries in the Pfam database. For protein domain analysis, the default settings of Pfam 7.0 [“global and local alignments merged” and “Pfam gathering threshold (GA)”] were used.











