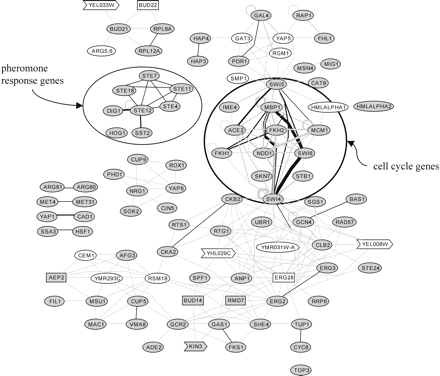
Visualization of the source gene pairs with highly similar target sets as a graph. Genes involved in the same biological processes are often connected and are thus close in the resulting graph; e.g., pheromone response genes or cell-cycle genes (encircled). Genes sharing highly similar target sets (P ≤ 10-12) are connected by gray edges or, if there is also a corresponding edge in one of the reference networks ppi2, mi3, and mips, by black edges. The thicker edges indicate that the respective source gene pair had significantly similar target sets in several network comparisons. White nodes: genes which are not present in any of the reference networks; these genes therefore are not adjacent to any confirmed edges. Gray nodes: genes present in at least one of the reference networks. Rectangular nodes: genes with unknown molecular function (BUD14, ERG28, RMD7, BUD22, and AEP2). Arrow-shaped nodes: genes of unknown biological process (KIN3, YEL008W, YEL033W, YHL029C).











