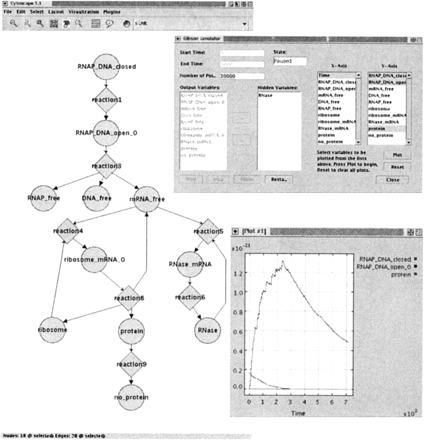
Cytoscape and the Systems Biology Workbench stochastically simulating gene regulation. In collaboration with the ERATO Systems Biology Workbench (SBW) project (Hucka et al. 2002), we developed a plug-in that allows Cytoscape to read and simulate SBML-encoded biochemical models. A Cytoscape network view of the SBML model is shown (left), accompanied by a user interface to the simulator (top, right) and an x-y plot of stochastic simulation results (bottom, right). In the x-y plot, the top curve shows the concentration of the N protein, which regulates genes expressed early in λ phage life cycle. Details are available at http://www.cytoscape.org/plugins/SBW/.











