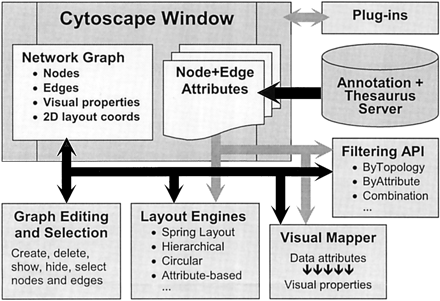
Figure 2
Schematic overview of the Cytoscape Core architecture. The Cytoscape window is the primary visual and programmatic interface to Cytoscape and contains the network graph and attribute data structures. Core methods that operate on these structures are graph editing, graph layout, attribute-to-visual mapping, and graph filtering. Annotations are available through a separate server.











