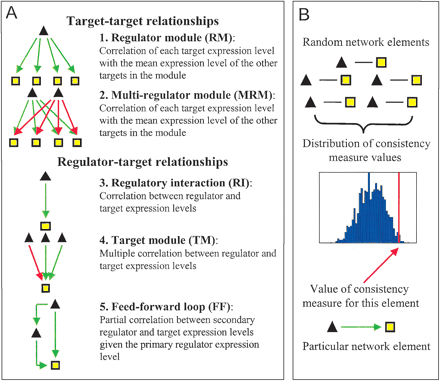
(A) Regulatory network elements or building blocks studied. The element types are classified into those involving only target-target relationships and those involving only regulator-target relationships. For each type of element, a diagram illustrating a typical element structure is shown. The black triangles are transcriptional regulators, and yellow boxes are transcription factor target genes. The red and green arrows correspond to repressing and activating regulatory interactions respectively. The consistency measure used to evaluate the agreement between a particular element and gene expression data is also indicated. Feed-forward loops are not considered to be a basic building block of the network, but are included here because ignoring them could lead to overestimating the consistency of regulatory interactions. (B) A schematic illustration of the strategy used to calculate the statistical significance of a particular value of a consistency measure (in this case Pearson correlation coefficient for a regulatory interaction). A set of random network elements corresponding to the particular network element studied is created, and the same consistency measure is calculated for each random element. The resulting distribution of values is used to evaluate the statistical significance of the true value of a consistency measure in the form of a P-value. The P-value is computed as the fraction of random elements with the squared consistency measure value higher than the squared observed value for the actual element. This corresponds to the estimated probability of observing a squared consistency measure value as high as the one for the actual element for a regulatory network element with random regulator and target genes.











