Limitations of Quantitative Gene Regulation Models: A Case Study
- Philip M. Kim1,2 and
- Bruce Tidor1,3,4,5
- 1 Computer Science and Artificial Intelligence Laboratory, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA
- 2 Department of Chemistry, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA
- 3 Biological Engineering Division, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA
- 4 Department of Electrical Engineering and Computer Science, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA
Abstract
Understanding the relationship between network structure and behavior is fundamental to the field of computational and systems biology. A particularly important distinction is the extent to which qualitative aspects of network performance are encoded in network topology as opposed to being determined through quantitative details, such as those of kinetics. Here, we develop a general and rigorous mathematical framework for the analysis of genetic networks and apply it to a family of synthetic gene networks. A key feature of our methodology involves determining network behavior that is insensitive to kinetic parameters such as rate constants and nonlinear functional dependencies of rates on molecular concentrations. Results indicate that behavior observed in some networks cannot be reconciled with standard gene expression and regulation models. We explore relaxing model assumptions to explain the observed behavior, allowing for both dynamicand stochastic phenomena, and propose an alternative model. Our alternative model includes the suggestion of a new mechanism by which the counterintuitive behavior could be achieved; central to the model is the assumption that the Clp protein degradation system, which is responsible for the regulatory proteins used in this study, becomes saturated.
Footnotes
-
Article and publication are at http://www.genome.org/cgi/doi/10.1101/gr.1207003.
-
↵6 We need the strict monotonic property of the degradation rates in order to be able to compute the inverses, degri-1 and degpi-1.
-
↵7 Guet et al. (2002) added aTc at 100 ng ml-1 and IPTG at 1 mM, which is known to be about 10-fold higher than the saturating concentration of each effector (Lutz and Bujard 1997). Note that our treatment becomes somewhat less general by assuming effector saturation.
-
↵8 We can apply simple arguments about monotonicity here. Because we know that tr(pe) > tr(p) and all functions in 17 are monotonic, we can deduce that f(pe) > f(p).
-
↵9 We have seen in eq. 18 that fe = f(pe)>f(p), therefore, also
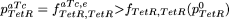 .
.
-
↵5 Corresponding author. E-MAIL tidor{at}mit.edu; FAX (617)252-1816.
-
- Accepted September 19, 2003.
- Received January 22, 2003.
- Cold Spring Harbor Laboratory Press











