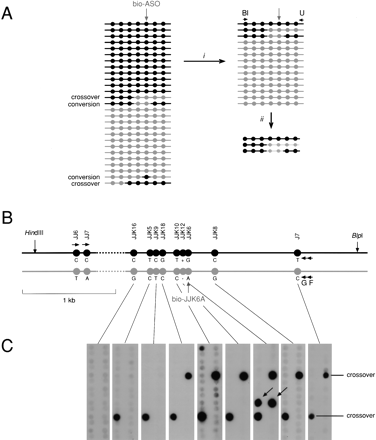
Use of DEASH to analyze recombination in human sperm. (A) The principle of detection. Sperm DNA molecules from a region with multiple SNP heterozygosities (circles) will contain a low frequency of recombinant molecules (crossovers and gene conversions). Sperm DNA fractionation (i) using a bio-ASO specific for the gray haplotype will recover gray haplotype molecules, plus any recombinant molecules that include the gray selector site, and will deplete though not totally eliminate molecules from the black haplotype. PCR using an allele-specific primer from the black haplotype (Bl) plus a universal (not allele-specific) primer (U) (ii) will generate PCR products only from recombinant molecules and from any remaining molecules of the black haplotype. (B) The HLA-DMB target region, as in Fig. 4, in the man tested. Sperm DNA (12 μg) was digested with HindIII plus BlpI and enriched using bio-JJK6A directed to the gray haplotype. Assays as in Fig. 4, as well as Poisson analyses of extreme dilutions of the DNA, showed that the enriched DNA contained 440,000 amplifiable molecules of the gray haplotype and was 130-fold enriched. Multiple aliquots of this DNA, each containing 4000 molecules of the gray haplotype plus 30 remaining molecules of the black haplotype, were amplified using the allele-specific primer JJ6C and the universal primer F, then re-amplified with primer JJ7C plus the universal primer G. (C) These re-amplified PCR products were dot-blot hybridized with 32P-labeled ASOs specific for each of the gray alleles. Four recombinants were detected, with hybridization signals consistent with them being present at a level of 3% of the PCR products from the remaining black haplotype. Two were crossovers with exchange points mapping to the intervals JJK9-JJK18 and JJK16-JJK5, respectively. The other two were conversions (arrowed) affecting only the selector site JJK6. These conversions were detected among 28 PCRs containing 112,000 molecules of the gray haplotype, giving a conversion rate of 2/112,000 = 1.8 × 10-5 per sperm.











