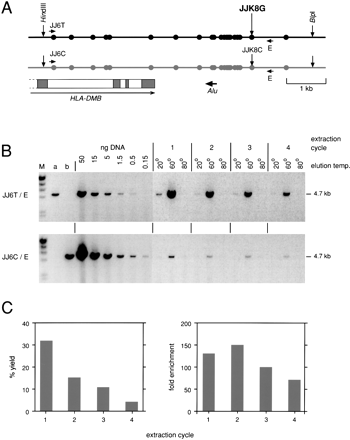
Bulk separation of haplotypes within the human MHC. (A) The target region containing the end of the HLA-DMB gene and multiple heterozygous SNPs (circles) in the individual tested (Jeffreys et al. 2001). The region was released on a 6.6-kb fragment by digesting sperm DNA with HindIII plus BlpI. The black haplotype was fractionated from 16μg DNA using DEASH directed to allele JJK8G, using four cycles of extraction. (B) Haplotype separation. Recovery of the black haplotype was assayed by 24 cycles of long PCR using the allele-specific primer JJ6T plus primer E and comparing PCR product yields from aliquots of the eluates (3% of each eluate tested) with yields from decreasing inputs of unfractionated digested DNA. Contamination with the gray haplotype was similarly assayed using primer JJ6C (27 cycles). Genomic DNA from individuals homozygous for JJ6T (a) or JJ6C (b) (5 ng per PCR) provided a control for allele-specificity; these primers were chosen for their high specificity, with each showing a >1000 fold discrimination between alleles (data not shown). M, λ DNA × HindIII markers. (C) Yield and degree of enrichment estimated for each cycle of extraction/re-extraction.











