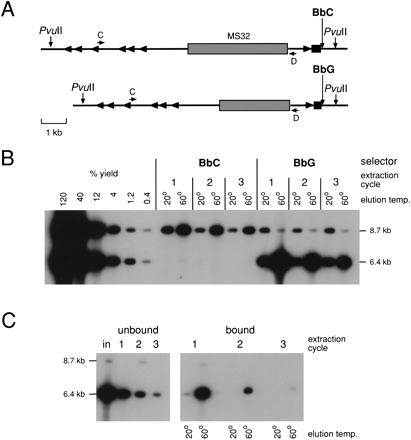
Separating haplotypes from human genomic DNA. (A) The target region, including the selector site (BbC/G) and minisatellite MS32 used to test for haplotype separation. Alu repeats are marked with arrowheads. (B) Haplotype separation. Four μg sperm DNA from a man with the two haplotypes shown in (A) was digested with PvuII and fractionated by DEASH using either bio-BbC or bio-BbG. Unbound DNA was subjected to two further cycles of extraction after adding bio-BbC or bio-BbG. Aliquots of eluted fractions (2% of each eluate) were assayed by long PCR amplification with primers C and D, followed by agarose gel electrophoresis and Southern blot hybridization with a 32P-labeled MS32 repeat probe. Yields were estimated by comparison with decreasing inputs of unfractionated PvuII-digested sperm DNA (48-0.16ng). The autoradiograph was overexposed to show traces of contaminating haplotype. (C) Assaying depletion of target from unbound DNA during DEASH. BbG-fractionated single-stranded DNA from B (60°C eluates pooled from all three extraction cycles) was subjected to three cycles of extraction/re-extraction using bio-BbG. Unbound (left) and bound (right) DNA recovered after each cycle was analyzed by PCR as in B and compared to input DNA (in). Note that primers C and D are located away from the selector site, allowing the unbound DNA to be analyzed by PCR even though it contains competitor BbC ASO. The faint residual 8.7-kb signal in the re-extracted fraction 1 is that expected for a single molecule of DNA, and given repeat instability and recombinational activity at MS32 (Jeffreys et al. 1998a,b), may be a mutant/recombinant molecule containing BbG rather than a contaminating BbC allele. For this reason, the MS32 system cannot be used to estimate levels of additional purification achieved by re-extraction.











