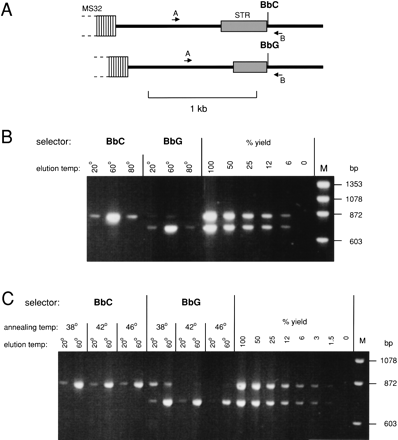
Use of DEASH to separate haplotypes from PCR products. (A) The target region adjacent to human minisatellite MS32 (Jeffreys et al. 1998b) in the person analyzed, including a heterozygous SNP (BbC/G) used for DEASH selection and a length-heterozygous STR locus. This region was PCR-amplified from blood DNA using primers A and B. (B) Haplotype separation using BbC/G. Ten pg PCR products were fractionated by DEASH using biotinylated BbC or BbG ASOs, with annealing at 42°C. Hybrids retained on Dynabeads were eluted sequentially at the indicated temperatures. Aliquots of eluted single-stranded DNA (4% of each eluate) were amplified using primers A and B, along with decreasing inputs of unfractionated PCR products (200-12 fg DNA) to estimate yield of recovered DNA, and analyzed by agarose gel electrophoresis followed by staining with ethidium bromide. M, marker DNA. (C) Effects of varying the annealing temperature on yield and purification.











