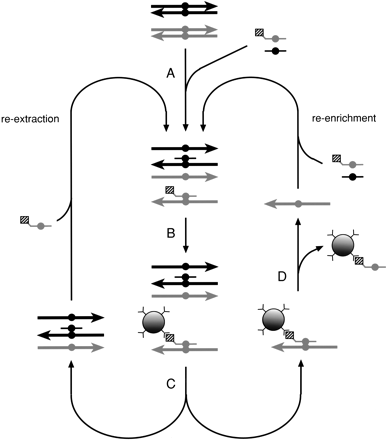
The principles of DEASH. (A) DNA containing a mixture of molecules differing by the chosen base substitution (circle) is mixed with a biotinylated ASO (gray, with biotin shown as a square) and the corresponding competitor ASO (black) directed to the alternative variant, then denatured and hybridized. (B) Streptavidin-coated paramagnetic beads (Dynal) are added to bind the bio-ASO/target hybrid and are then separated with a magnet (C). Bound hybrid is thermally eluted (D), releasing the enriched single-stranded target and leaving the bio-ASO still bound to the beads. Further target can be recovered by adding bio-ASO to the unbound DNA, denaturing, and processing as above (re-extraction). Enriched target can be purified further by adding buffer, bio-ASO, and competitor ASO, followed by DEASH (re-enrichment).











