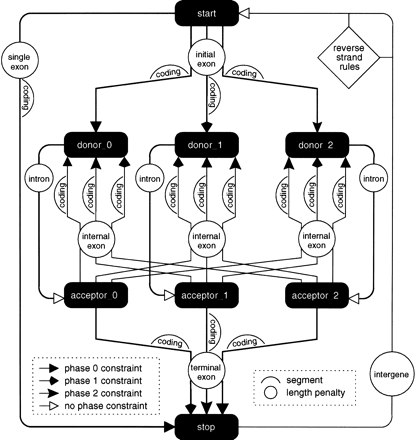
A pictorial representation of a GAZE-XML model for multiple genes on both strands. The features are represented by filled boxes, and ’source → target' rules by different types of arrows, each corresponding to a phase constraint as explained in the text. The labeled circles give the name of the length penalty function used for each pair of features, which are themselves defined elsewhere in the configuration file (not shown); the labeled humps indicate the segments that contribute to the score for each pair of features, where “coding” humps are the likely_coding segments referred to in the text. The rules for reverse-strand target features are not shown in their entirety, for clarity, but are formed by a simple reverse complementation of the forward-strand rules. Also omitted are the BEGIN and END features (which mark the two ends of the sequence being searched for genes, and act respectively as source and target to every other feature), as well as the distance, interruption, and DNA constraints explained in the text. The XML configuration file contains a directive to create three separate features for each predicted splice site seen in the GFF file. The effect of this, together with phase constraints between pairs of features giving rise to exons, is to carry forward whether each intron interrupts a codon at position 0, 1, or 2 to the rest of the gene structure, allowing us to ensure that the length of the coding part of each predicted gene is divisible by three.











