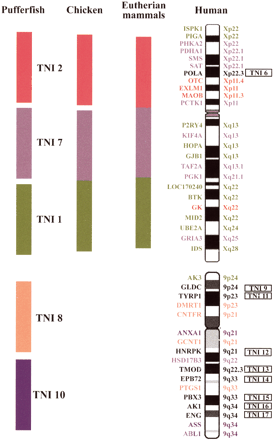
Syntenic relationships of human chromosomes X and 9 with theTetraodon genome and an evolutionary model for the mammalian X. The top shows the common phylogenetic origin of pufferfish chromosomes TNI 1, TNI 2, and TNI 7 and the human X. At left, the three Tetraodon blocks of conserved synteny are depicted as green (TNI 1), red (TNI 2), and purple (TNI 7) bars. Atright is shown the mosaic synteny of human X and the chromosomal localizations of the comparatively mapped genes. The gene order is based on genomic sequence data (http://www.genome.ucsc.edu/cgi-bin/hgGateway December 2001). ThePOLA gene, indicated in black, maps to a differentTetraodon chromosome (TNI 6) and, thus, does not belong to the delineated synteny groups. The two central schematic drawings represent hypothetical ancestral X chromosomes. Because human X orthologous genes from TNI 1 and TNI 7 are linked in chicken, we conclude that an ancestral X autosome contained these two sets of Xp and Xq genes even before the avian-mammalian split. Because the human X orthologs belonging to the TNI 2 block are autosomal in marsupials and monotremes, they represent a recent addition to the eutherian sex chromosomes. For simplicity, the three X syntenic Tetraodonchromosomes are depicted as blocks on the hypothetical X ancestors. However, this does not reflect the real gene order, which may differ significantly from both the order in the three Tetraodonchromosomes and in the human X. At bottom is shown a much lower degree of synteny conservation for human chromosome 9 in the pufferfish than for the X. Comparative mapping of human 9 orthologous genes reveals two syntenic blocks on TNI 8 (orange) and TNI 10 (blue). One gene each reside on TNI 1 (green) and TNI 7 (purple), respectively. The genes indicated in black all map to different TNI chromosomes.











